Achondroplasia & FGFR3
This page was produced as an assignment for Genetics 677 at UW-Madison, Spring 2010
In addition to providing the gene and protein accession information for all of the homologs of FGFR3 that I could find on Homologene, I attempted to find images of as many achondroplastic phenotypes as I could. Interestingly there are breeds of animals in which people have selectively bred for this mutation, and I have noted this under the images. Additionally, images of this mutant phenotype could only be found in the mammalian homologs, which could indicate that a mammal would probably the best model organism for any studies about the mutation and it's effects on bone development and visible phenotype.
Homologs of FGFR3 Mutant Phenotype

Dachshunds are bred to have achondroplasia.
Canis familiaris
Gene Name: FGFR3
Gene Accession Number: XM_545926.2
Protein Accession Number: XP_545926.2

Irish Dexter is an achondroplastic cattle breed
Bos taurus
Gene Name: FGFR3
Gene Accession Number: NM_174318.3
Protein Accession Number: NP_776743.1
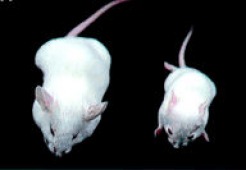
A mutant mouse (right) compared with a wild type
Rattus norvegicus
Gene Name: FGFR3
Gene Accession Number: NM_053429.1
Protein Accession Number: 445881.1
Gene Name: FGFR3
Gene Accession Number: NM_053429.1
Protein Accession Number: 445881.1
Gallus gallus
Gene Name: FGFR3
Gene Accession Number: NM_205509.1
Protein Accession Number: NP_990840.1
Gene Name: FGFR3
Gene Accession Number: NM_205509.1
Protein Accession Number: NP_990840.1

The 'bag of worms' phenotype that often results from egl
Caenorhabditis elegans
Gene Name: egl-15 (egg-laying defect 15)
Gene Accession Number: NM_001029554.2
Protein Accession Number: 001024725.1
Gene Name: egl-15 (egg-laying defect 15)
Gene Accession Number: NM_001029554.2
Protein Accession Number: 001024725.1
BLAST search
A BLAST search on the DNA sequence was run on the NCBI website. All of the homologs mentioned above, except for C.elegans (unfortunately its homolog did not come up in the search) had BLAST E-values of 0.0. An E-value of 1 implies that the sequence came up in the search merely by chance, as the value decreases towards 0, the match is less and less likely to be due to chance. So, the search provided very useful alignments. Additionally, most of the sequences had high alignment scores, as can be seen below:

