Achondroplasia & FGFR3
This page was produced as an assignment for Genetics 677 at UW-Madison, Spring 2010
DNA Motifs
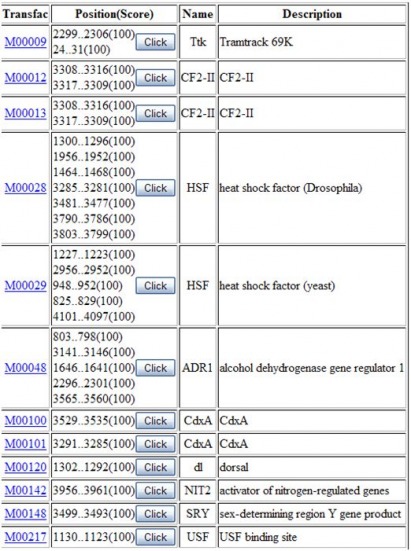
To explore the motifs within the FGFR3 gene, the FASTA sequence was processed with Motif Finder. Initially, I just used the supplied parameters with a cut-off score of 85. This produced 118 motifs, which seemed like a lot. After trying several different values for the cut-off score, I realized that I had to be very stringent and use a score of 100. This still produced 12 motifs.
However, if you look at the size of these motifs they are very small, often 5-7 bp in length. So even though they match perfectly, the size of the motif indicates that this would be a pretty easy size to match. This suspicion that these motifs are not very accurate are supported by the fact that none of them really seem to relate to the function of my gene.
Next, I tried using Meme to investigate my sequence. I entered in that it should come up with a maximum of 20 motifs, and it came up with all 20. This large number of motifs is similar to what happened on motif finder, which leads me to question how accurate these are, and what information about my gene I can accurately infer from them.
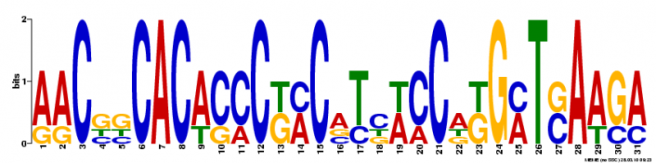
However, I was much more successful in finding motifs of larger sizes, which makes me think that they are more likely to have something to do with the function of my gene. I also really like the way Meme aids in visualizing the sequence. I have chosen two of the larger motifs and shown them below.
Next, I tried using Meme to investigate my sequence. I entered in that it should come up with a maximum of 20 motifs, and it came up with all 20. This large number of motifs is similar to what happened on motif finder, which leads me to question how accurate these are, and what information about my gene I can accurately infer from them.
However, I was much more successful in finding motifs of larger sizes, which makes me think that they are more likely to have something to do with the function of my gene. I also really like the way Meme aids in visualizing the sequence. I have chosen two of the larger motifs and shown them below.
This came up as the first motif and covers 36 sites. It also occurs five times towards the end of the gene sequence.
Another relatively large motif with 31 sites. This motif can be found a total of four times within the gene.