Achondroplasia & FGFR3
This page was produced as an assignment for Genetics 677 at UW-Madison, Spring 2010
Phylogentic Trees
Using the homologs shown on the homology page, the mRNA FASTA sequences were entered into Phylogeny.fr using MUSCLE alignment, Maximum likelihood, and Treedyn to construct the tree.
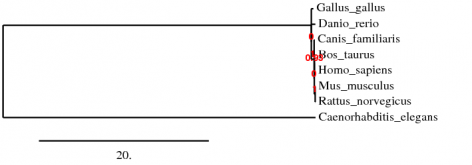
The results:
The results:
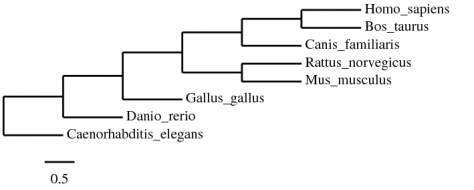
The same tree shown as a cladogram to better see the relationships:
The results from these tree seem to agree with evolutionary relationships. C. elegans is the most removed, then G. gallus, and D. rerio. The mammals are all closely grouped, and showing that M. musculus and R. norvegicus are closely related, which makes sense and supports the tree.
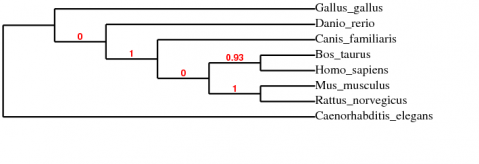
Next, I tried the "a la carte" mode with MUSCLE alignment, and Treedyn drawing, but using parsimony to calculate the relationships. It produced slightly different results:
Next, I tried the "a la carte" mode with MUSCLE alignment, and Treedyn drawing, but using parsimony to calculate the relationships. It produced slightly different results:
In this tree, D. rerio branches off before G. gallus, which does seem to make more sense evolutionarily. It is interesting that such a different result could be obtained by changing the statistical method, and supports the idea that one should calculate a phylogenetic tree in several different ways to ensure that the conclusions are consistent. See the FGFR3 protein phylogeny page for more comparisons.