Achondroplasia & FGFR3
This page was produced as an assignment for Genetics 677 at UW-Madison, Spring 2010
Protein Domains
The domains of the FGFR3 protein were found using SMART and drawn using MyDomains. This protein is characterized by three segments: the extracellular portion which binds the ligands, the transmembrane portion, and the tyrosine kinase which acts to phosphorylate other proteins and such within the cell to start a signaling pathway.
The tyrosine kinase is a subfamily of protein kinases. This is a domain found in many proteins and in many species from single celled organisms to humans, performing a multitude of functions within the cell. [1] Its basic mode of action is to transfer a phosphate group to a target protein, thereby causing a conformational change in the target which affects its function. The tyrosine kinase is shown below in orange and occupies a large part of the protein, from Amino acid 472 to 748.
The mutation that causes achondroplasia is located within the transmembrane domain, at position 380, highlighted on the domain map with a little red diamond. The amino acid sequence of the transmembrane region, as determined by SMART, is as follows:
YAGILSYGVGFFLFILVVAAVTL
All of these amino acids are hydrophobic, some of them extremely so, which makes it easy for them to reside within the lipid bilayer. [2] The mutation however, changes 380 glycine to an arginine, which only has a slightly hydrophobic side chain, while the rest of the amino acid is not. Further investigation into how this shift in the properties of the amino acids composing this portion of the protein affects its function could provide useful.
The tyrosine kinase is a subfamily of protein kinases. This is a domain found in many proteins and in many species from single celled organisms to humans, performing a multitude of functions within the cell. [1] Its basic mode of action is to transfer a phosphate group to a target protein, thereby causing a conformational change in the target which affects its function. The tyrosine kinase is shown below in orange and occupies a large part of the protein, from Amino acid 472 to 748.
The mutation that causes achondroplasia is located within the transmembrane domain, at position 380, highlighted on the domain map with a little red diamond. The amino acid sequence of the transmembrane region, as determined by SMART, is as follows:
YAGILSYGVGFFLFILVVAAVTL
All of these amino acids are hydrophobic, some of them extremely so, which makes it easy for them to reside within the lipid bilayer. [2] The mutation however, changes 380 glycine to an arginine, which only has a slightly hydrophobic side chain, while the rest of the amino acid is not. Further investigation into how this shift in the properties of the amino acids composing this portion of the protein affects its function could provide useful.
The extracellular portion of the protein is made up of three IG domains of which two (in green) are part of the subfamily specifically found in FGFR receptors. In general, IG domains can perform a number of functions including cell-to-cell recognition, muscle development, and cell surface receptor which is how it is used in this protein. [3]
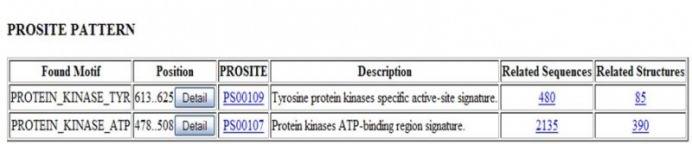
In addition to mapping the domains, I ran the protein FASTA sequence through Motif Finder and came up with two motifs:
In addition to mapping the domains, I ran the protein FASTA sequence through Motif Finder and came up with two motifs:
These results seem consistent with the other findings that this protein has a tyrosine protein kinase.
References
[1] Manning G., Plowman GD, Hunter T, Sudarsanam S. (2002). Evolution of protein kinase signaling from yeast to man. Trends Biochem Sci. Oct;27(10):514-20.
[2] M.J. Betts, R.B. Russell. Amino acid properties and consequences of subsitutions.
In Bioinformatics for Geneticists, M.R. Barnes, I.C. Gray eds, Wiley, 2003. http://www.russell.embl.de/aaTeis/hydrophobic.html
[3] Teichmann SA, Chothia C. (2000). Immunoglobulin superfamily proteins in Caenorhabditis elegans. J Molecular Biology. Mar 10;296(5):1367-83.
[2] M.J. Betts, R.B. Russell. Amino acid properties and consequences of subsitutions.
In Bioinformatics for Geneticists, M.R. Barnes, I.C. Gray eds, Wiley, 2003. http://www.russell.embl.de/aaTeis/hydrophobic.html
[3] Teichmann SA, Chothia C. (2000). Immunoglobulin superfamily proteins in Caenorhabditis elegans. J Molecular Biology. Mar 10;296(5):1367-83.