Achondroplasia & FGFR3
This page was produced as an assignment for Genetics 677 at UW-Madison, Spring 2010
Protein Phylogeny
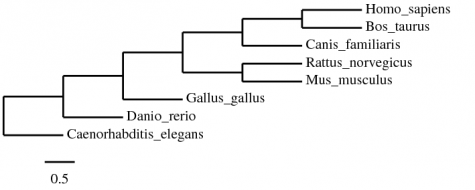
To further explore the phylogeny of FGFR3, the protein FASTA sequence was entered at Phylogeny.fr, using the "a la carte" mode. For the first try ClustalW alignment, maximum likelihood statistics, and Treedyn were used, just like when constructing the mRNA phylogeny trees. Surprisingly, this did not produce the same tree:
This tree differs from the mRNA one derived from the same a la carte options in that D. rerio branches off before G. gallus. This result, however makes more sense from an evolutionary standpoint.
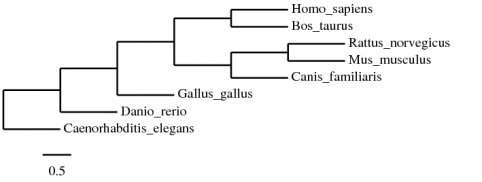
For another tree to compare these contrasting results with, I ran the protein sequence again but choosing parsimony for the calculations:
For another tree to compare these contrasting results with, I ran the protein sequence again but choosing parsimony for the calculations:
This tree helps to support the other tree regarding when D. rerio and G. gallus should branch off. However, other relationships within the mammals have changed. This tree shows C. familiaris as being more closely related to M. musculus than H. sapiens, unlike both the other protein-derived tree as well as the mRNA-derived tree that used parsimony.
The differences observed in these trees demonstrate how much variation can occur in phylogenetics, and how much both the methods and the kind of sequences used can alter the conclusions.
The differences observed in these trees demonstrate how much variation can occur in phylogenetics, and how much both the methods and the kind of sequences used can alter the conclusions.